This page was created for University of Wisconsin undergraduate course Genetics 564
What is Phylogeny?
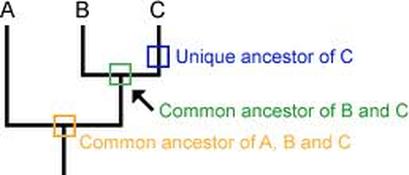 Figure 1: Depiction of branching in the Tree of Life
Figure 1: Depiction of branching in the Tree of Life
Phylogeny is the study of the tree of life- the ancestral history all species share. As time went on, DNA mutations caused certain organisms to have differences from others, leading to branching in the evolutionary tree. These differences accumulated causing more and more forks in the road until we reached the species we have today. This process of branch development is demonstrated in Figure 1. The oldest common ancestor begins at the bottom of the image. The yellow square, or the 'node', marks where a mutation occurred in a certain organism, causing a fork in evolution. The green node marks a divergence in characteristics between species B and C. Thus B and C in this image share a more recent common ancestor than with A and are more closely related.
EFHC1 Protein Phylogeny
To examine the common ancestors of the EFHC1 protein, homologs were confirmed using homologene, a database of known protein matches among species. The sequences of amino acids in the proteins of each specie were examined side by side and aligned, looking for common sequences using ClustalOmega. An example of the Clustal Omega results is shown in image 1, which highlights in a column in one color when species share the same amino acid in a certain position.
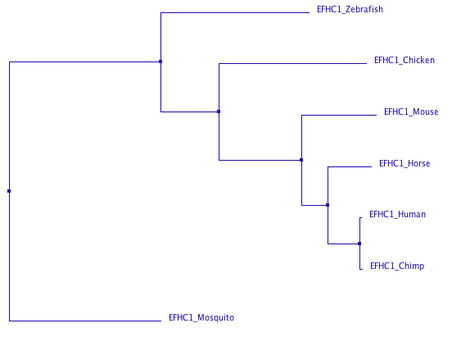
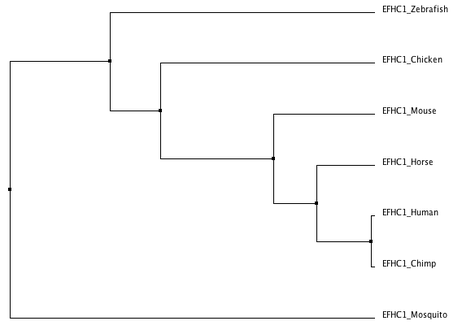
This analysis was then used to create phylogenetic trees of the progression of the protein through evolutionary time still using the ClustalOmega program. Trees were created using neighbor joining by percent identity (image 2, left) and average distance by percent identity (image 3, right). A percent identity tree is formed by comparing the number of identical bases per 100 base pairs in species' sequences. The neighbor joining method attempts to find the shortest possible branch lengths, while the average distance method joins taxa by studying the genetic distances between sequences and creating clusters of species with the least amount of dissimilarities.
Analysis
As is clearly shown, both methods of tree modeling agreed on the evolutionary progression of EFHC1. Both models placed mosquitos as the outgroup and most distantly related, and placed chimps as human EFHC1's most closely related relative. These evolutionary trees coincide with the accepted evolution of species as well, as chimpanzees are humans closest related specie. As you move backwards through evolutionary time, humans are then most related to other mammals and vertebrates, finally being least related to an insect, the mosquito. Thus I find the evolution of EFHC1 protein to be accurate from these phylogenic trees.
References:
Figure 1 : Understanding Evolution. http://evolution.berkeley.edu/evolibrary/article/evo_05